Partial Least Squares in R
06.19.2021
Intro
Partial Least Squares is a machine learning model that helps solbe issues with multicollinearity. It has advantages of PCA regression in the sense that it is still easily interpretable and has good performance. In this article, we will learn how to use partial least squares in R.
Data
For this tutorial, we will use the Boston data set which includes
housing data with features of the houses and their prices. We would like
to predict the medv column or the medium value.
library(MASS)
data(Boston)
str(Boston)## 'data.frame': 506 obs. of 14 variables:
## $ crim : num 0.00632 0.02731 0.02729 0.03237 0.06905 ...
## $ zn : num 18 0 0 0 0 0 12.5 12.5 12.5 12.5 ...
## $ indus : num 2.31 7.07 7.07 2.18 2.18 2.18 7.87 7.87 7.87 7.87 ...
## $ chas : int 0 0 0 0 0 0 0 0 0 0 ...
## $ nox : num 0.538 0.469 0.469 0.458 0.458 0.458 0.524 0.524 0.524 0.524 ...
## $ rm : num 6.58 6.42 7.18 7 7.15 ...
## $ age : num 65.2 78.9 61.1 45.8 54.2 58.7 66.6 96.1 100 85.9 ...
## $ dis : num 4.09 4.97 4.97 6.06 6.06 ...
## $ rad : int 1 2 2 3 3 3 5 5 5 5 ...
## $ tax : num 296 242 242 222 222 222 311 311 311 311 ...
## $ ptratio: num 15.3 17.8 17.8 18.7 18.7 18.7 15.2 15.2 15.2 15.2 ...
## $ black : num 397 397 393 395 397 ...
## $ lstat : num 4.98 9.14 4.03 2.94 5.33 ...
## $ medv : num 24 21.6 34.7 33.4 36.2 28.7 22.9 27.1 16.5 18.9 ...Basic Partial Least Squares in R
To build a Partial Least Squares model, we can use the plsr method
from the pls package. We pass two parameters, the model equation which
says, medv ~ ., predict medium value by all other predictors, and our
Boston data set.
library(pls)## Warning: package 'pls' was built under R version 4.0.5
##
## Attaching package: 'pls'
## The following object is masked from 'package:stats':
##
## loadingsmodel <- plsr(medv ~ ., data = Boston)
summary(model)## Data: X dimension: 506 13
## Y dimension: 506 1
## Fit method: kernelpls
## Number of components considered: 13
## TRAINING: % variance explained
## 1 comps 2 comps 3 comps 4 comps 5 comps 6 comps 7 comps 8 comps
## X 80.51 94.45 98.97 99.34 99.80 99.91 99.94 99.97
## medv 24.23 26.94 32.05 51.05 60.08 62.49 66.54 68.31
## 9 comps 10 comps 11 comps 12 comps 13 comps
## X 99.99 100.00 100.00 100.00 100.00
## medv 69.03 71.07 72.75 73.08 74.06Modeling Partial Least Squares in R with Caret
We will now see how to model a lasso regression using the Caret
package. We will use this library as it provides us with many features
for real life modeling.
To do this, we use the train method. We pass the same parameters as
above, but in addition we pass the method = 'lasso' model to tell
Caret to use a lasso model.
set.seed(1)
library(caret)## Loading required package: lattice
## Loading required package: ggplot2
##
## Attaching package: 'caret'
## The following object is masked from 'package:pls':
##
## R2model <- train(
medv ~ .,
data = Boston,
method = 'pls'
)
model## Partial Least Squares
##
## 506 samples
## 13 predictor
##
## No pre-processing
## Resampling: Bootstrapped (25 reps)
## Summary of sample sizes: 506, 506, 506, 506, 506, 506, ...
## Resampling results across tuning parameters:
##
## ncomp RMSE Rsquared MAE
## 1 7.971850 0.2530128 5.782578
## 2 7.851798 0.2752155 5.665435
## 3 7.574906 0.3247404 5.280531
##
## RMSE was used to select the optimal model using the smallest value.
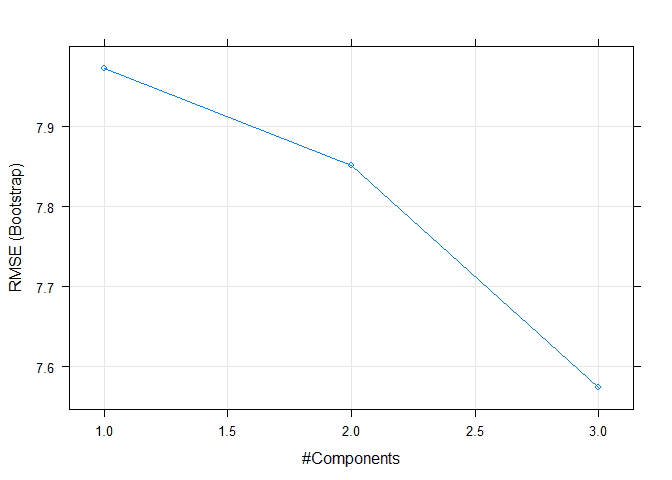
## The final value used for the model was ncomp = 3.Here we can see that caret automatically trained over multiple hyper parameters. We can easily plot those to visualize.
plot(model)Preprocessing with Caret
One feature that we use from Caret is preprocessing. Often in real life
data science we want to run some pre processing before modeling. We will
center and scale our data by passing the following to the train method:
preProcess = c("center", "scale").
set.seed(1)
library(caret)
model2 <- train(
medv ~ .,
data = Boston,
method = 'pls',
preProcess = c("center", "scale")
)
model2## Partial Least Squares
##
## 506 samples
## 13 predictor
##
## Pre-processing: centered (13), scaled (13)
## Resampling: Bootstrapped (25 reps)
## Summary of sample sizes: 506, 506, 506, 506, 506, 506, ...
## Resampling results across tuning parameters:
##
## ncomp RMSE Rsquared MAE
## 1 6.461016 0.5070922 4.422147
## 2 5.009639 0.7015362 3.427887
## 3 4.995151 0.7047935 3.419506
##
## RMSE was used to select the optimal model using the smallest value.
## The final value used for the model was ncomp = 3.plot(varImp(model2))Splitting the Data Set
Often when we are modeling, we want to split our data into a train and
test set. This way, we can check for overfitting. We can use the
createDataPartition method to do this. In this example, we use the
target medv to split into an 80/20 split, p = .80.
This function will return indexes that contains 80% of the data that we should use for training. We then use the indexes to get our training data from the data set.
set.seed(1)
inTraining <- createDataPartition(Boston$medv, p = .80, list = FALSE)
training <- Boston[inTraining,]
testing <- Boston[-inTraining,]We can then fit our model again using only the training data.
set.seed(1)
model3 <- train(
medv ~ .,
data = training,
method = 'pls',
preProcess = c("center", "scale")
)
model3## Partial Least Squares
##
## 407 samples
## 13 predictor
##
## Pre-processing: centered (13), scaled (13)
## Resampling: Bootstrapped (25 reps)
## Summary of sample sizes: 407, 407, 407, 407, 407, 407, ...
## Resampling results across tuning parameters:
##
## ncomp RMSE Rsquared MAE
## 1 6.493166 0.4973332 4.512294
## 2 5.047896 0.6977929 3.472897
## 3 5.025694 0.7015002 3.463829
##
## RMSE was used to select the optimal model using the smallest value.
## The final value used for the model was ncomp = 3.Now, we want to check our data on the test set. We can use the subset
method to get the features and test target. We then use the predict
method passing in our model from above and the test features.
Finally, we calculate the RMSE and r2 to compare to the model above.
test.features = subset(testing, select=-c(medv))
test.target = subset(testing, select=medv)[,1]
predictions = predict(model3, newdata = test.features)
# RMSE
sqrt(mean((test.target - predictions)^2))## [1] 5.247168# R2
cor(test.target, predictions) ^ 2## [1] 0.6968392Cross Validation
In practice, we don’t normal build our data in on training set. It is
common to use a data partitioning strategy like k-fold cross-validation
that resamples and splits our data many times. We then train the model
on these samples and pick the best model. Caret makes this easy with the
trainControl method.
We will use 10-fold cross-validation in this tutorial. To do this we
need to pass three parameters method = "repeatedcv", number = 10
(for 10-fold). We store this result in a variable.
set.seed(1)
ctrl <- trainControl(
method = "cv",
number = 10,
)Now, we can retrain our model and pass the trainControl response to
the trControl parameter. Notice the our call has added
trControl = set.seed.
set.seed(1)
model4 <- train(
medv ~ .,
data = training,
method = 'pls',
preProcess = c("center", "scale"),
trControl = ctrl
)
model4## Partial Least Squares
##
## 407 samples
## 13 predictor
##
## Pre-processing: centered (13), scaled (13)
## Resampling: Cross-Validated (10 fold)
## Summary of sample sizes: 367, 366, 367, 366, 365, 367, ...
## Resampling results across tuning parameters:
##
## ncomp RMSE Rsquared MAE
## 1 6.379015 0.5273565 4.499162
## 2 4.869466 0.7210883 3.498225
## 3 4.810612 0.7315127 3.420090
##
## RMSE was used to select the optimal model using the smallest value.
## The final value used for the model was ncomp = 3.This results seemed to have improved our accuracy for our training data. Let’s check this on the test data to see the results.
test.features = subset(testing, select=-c(medv))
test.target = subset(testing, select=medv)[,1]
predictions = predict(model4, newdata = test.features)
# RMSE
sqrt(mean((test.target - predictions)^2))## [1] 5.247168# R2
cor(test.target, predictions) ^ 2## [1] 0.6968392Tuning Hyper Parameters
We can use caret to tune our models by using the tuneGride parameter.
For pls we want to tune the number of components used. In this
example, we tune of 1 to 10 components.
set.seed(1)
tuneGrid <- expand.grid(
ncomp = seq(1, 10, by = 1)
)
model5 <- train(
medv ~ .,
data = training,
method = 'pls',
preProcess = c("center", "scale"),
trControl = ctrl,
tuneGrid = tuneGrid
)
model5## Partial Least Squares
##
## 407 samples
## 13 predictor
##
## Pre-processing: centered (13), scaled (13)
## Resampling: Cross-Validated (10 fold)
## Summary of sample sizes: 367, 366, 367, 366, 365, 367, ...
## Resampling results across tuning parameters:
##
## ncomp RMSE Rsquared MAE
## 1 6.379015 0.5273565 4.499162
## 2 4.869466 0.7210883 3.498225
## 3 4.810612 0.7315127 3.420090
## 4 4.810029 0.7313804 3.465214
## 5 4.785486 0.7352180 3.458035
## 6 4.755067 0.7376506 3.446745
## 7 4.736177 0.7395488 3.430267
## 8 4.739281 0.7397892 3.427183
## 9 4.736283 0.7401848 3.427627
## 10 4.738701 0.7399414 3.427894
##
## RMSE was used to select the optimal model using the smallest value.
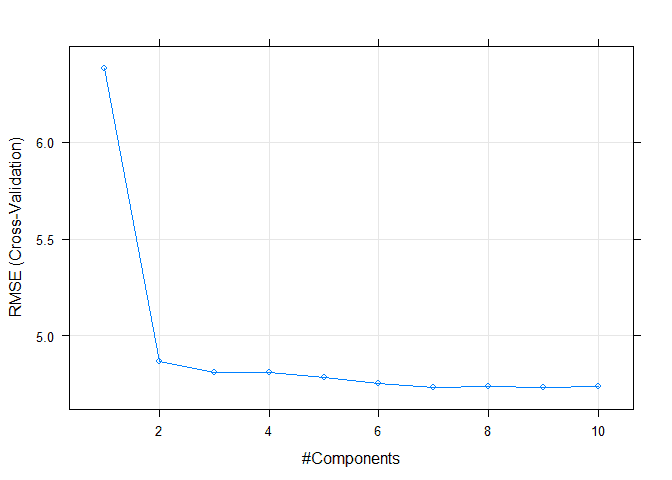
## The final value used for the model was ncomp = 7.We can plot our tuning results afterwards to visualize the progress.
plot(model5)